Course Outline
-
segmentGetting Started (Don't Skip This Part)
-
segmentStatistics and Data Science: A Modeling Approach
-
segmentPART I: EXPLORING VARIATION
-
segmentChapter 1 - Welcome to Statistics: A Modeling Approach
-
segmentChapter 2 - Understanding Data
-
segmentChapter 3 - Examining Distributions
-
segmentChapter 4 - Explaining Variation
-
segmentPART II: MODELING VARIATION
-
segmentChapter 5 - A Simple Model
-
segmentChapter 6 - Quantifying Error
-
segmentChapter 7 - Adding an Explanatory Variable to the Model
-
7.4 Examining Residuals from the Model
-
segmentChapter 8 - Models with a Quantitative Explanatory Variable
-
segmentPART III: EVALUATING MODELS
-
segmentChapter 9 - Distributions of Estimates
-
segmentChapter 10 - Confidence Intervals and Their Uses
-
segmentChapter 11 - Model Comparison with the F Ratio
-
segmentChapter 12 - What You Have Learned
-
segmentFinishing Up (Don't Skip This Part!)
-
segmentResources
list full book
7.4 Examining Residuals From the Model
As we have said before, when we use a model to make predictions, our predictions are usually wrong. The residuals, or the distance of each observed score from the predicted score, give us an indication of how wrong our predictions are for each person.
Calculating Residuals From the Model
We calculated residuals from the empty model by subtracting the mean from each score. We use the same approach for the more complex model.
Using the model, we assign a predicted score to each observation. This time, however, the predicted score is not the Grand Mean for everyone, but one mean for females and another for males.
Notice that we use the same strategy to quantify the leftover error from any model. We can do the subtractions in R, just as we did for the empty model. We start with the TinyFingers data frame looking like this.
Sex Thumb Sex.predicted
1 female 56 59
2 female 60 59
3 female 61 59
4 male 63 65
5 male 64 65
6 male 68 65We then calculate the residual for each person by this subtraction: their observed score minus their score predicted by the model. We will save the result in a new variable in TinyFingers that we will call Sex.resid.
TinyFingers$Sex.resid <- TinyFingers$Thumb - TinyFingers$Sex.predicted
TinyFingers Sex Thumb Sex.predicted Sex.resid
1 female 56 59 -3
2 female 60 59 1
3 female 61 59 2
4 male 63 65 -2
5 male 64 65 -1
6 male 68 65 3Another, slightly easier, way to do the same thing is to use the resid() function, using the TinySex.model as the argument. Let’s try that, and save the results in a new variable, Sex.resid2. Then let’s print the updated version of TinyFingers.
TinyFingers$Sex.resid2 <- resid(TinySex.model)
TinyFingers Sex Thumb Sex.predicted Sex.resid Sex.resid2
1 female 56 59 -3 -3
2 female 60 59 1 1
3 female 61 59 2 2
4 male 63 65 -2 -2
5 male 64 65 -1 -1
6 male 68 65 3 3 Compare the two variables—Sex.resid and Sex.resid2—in the output. Notice that we get the same values for Sex.resid2 as we did for Sex.resid.
Finally, let’s compare the residuals from the TinySex.model to those from the TinyEmpty.model. Let’s first add a variable called Empty.pred using the predict() function, and print out TinyFingers again. (We could have called the variable anything; the point is to name it something meaningful—so in this case we called it Empty.pred, short for predicted from the empty model.)
TinyFingers$Empty.pred <- predict(TinyEmpty.model)
TinyFingers Sex Thumb Sex.predicted Sex.resid Sex.resid2 Empty.pred
1 female 56 59 -3 -3 62
2 female 60 59 1 1 62
3 female 61 59 2 2 62
4 male 63 65 -2 -2 62
5 male 64 65 -1 -1 62
6 male 68 65 3 3 62 Now use the resid() function to create a new variable in the TinyFingers data frame, Empty.resid. And print out the updated version of TinyFingers.
require(tidyverse)
require(mosaic)
require(Lock5Data)
require(supernova)
TinyFingers <- data.frame(
Sex = as.factor(rep(c("female", "male"), each = 3)),
Thumb = c(56, 60, 61, 63, 64, 68)
)
TinyEmpty.model <- lm(Thumb ~ NULL, data = TinyFingers)
TinySex.model <- lm(Thumb ~ Sex, data = TinyFingers)
TinyFingers <- TinyFingers %>% mutate(
Sex.predicted = predict(TinySex.model),
Sex.resid = Thumb - Sex.predicted,
Sex.resid2 = resid(TinySex.model),
Empty.pred = predict(TinyEmpty.model)
)
# generate residuals from TinyEmpty.model
TinyFingers$Empty.resid <-
# write code to print TinyFingers
TinyFingers$Empty.resid <- resid(TinyEmpty.model)
TinyFingers
ex() %>% {
check_object(., "TinyFingers")
check_output_expr(., "print(TinyFingers)")
}
Sex Thumb Sex.predicted Sex.resid Sex.resid2 Empty.pred Empty.resid
1 female 56 59 -3 -3 62 -6
2 female 60 59 1 1 62 -2
3 female 61 59 2 2 62 -1
4 male 63 65 -2 -2 62 1
5 male 64 65 -1 -1 62 2
6 male 68 65 3 3 62 6Graphing Residuals From the Model
You might wonder, why are we bothering to generate and save residuals? We will have a lot more to say about this later. But the short answer is: it helps us to understand the error around our model, and often suggests ways of improving the model.
Just as the first thing we do when looking at a data set is to examine the distributions of the variables, it is good to get in the habit of examining the distributions of residuals after we fit a new model.
Let’s go back to the full Fingers data frame. We fit the model lm(Thumb ~ Sex, data = Fingers) and saved the model in Sex.model. Using the resid() function, write some code to generate a new column in Fingers called Sex.resid (the residuals from the Sex.model).
require(tidyverse)
require(mosaic)
require(Lock5Data)
require(supernova)
Sex.model <- lm(Fingers$Thumb ~ Fingers$Sex)
# store the residuals from Sex.model
Fingers$Sex.resid <-
# This prints the first 10 rows of Fingers
head(select(Fingers, Thumb, Sex.resid), 10)
# store the residuals from Sex.model
Fingers$Sex.resid <- resid(Sex.model)
# This prints the first 10 rows of Fingers
head(select(Fingers, Thumb, Sex.resid), 10)
ex() %>% {
check_object(., "Fingers") %>%
check_column("Sex.resid") %>%
check_equal()
}
resid() to get the residuals for each case.In the following window, we have provided the code to create density histograms of Thumb in a facet grid by Sex. Try modifying it to generate density histograms of Sex.resid in a facet grid by Sex. Compare the histograms of residuals from the Sex.model with histograms of thumb length.
require(tidyverse)
require(mosaic)
require(Lock5Data)
require(supernova)
Sex.model <- lm(Fingers$Thumb ~ Fingers$Sex)
Fingers$Sex.resid <- resid(Sex.model)
# This creates histograms of Thumb for each Sex
# Modify it to create histograms of Sex.resid for each Sex
gf_dhistogram(~Thumb, data = Fingers) %>%
gf_facet_grid(Sex ~ .)
gf_dhistogram(~Sex.resid, data = Fingers) %>%
gf_facet_grid(Sex ~ .)
ex() %>% check_or(
check_function(., "gf_dhistogram") %>% {
check_arg(., "object") %>% check_equal()
check_arg(., "data") %>% check_equal()
},
override_solution(., "gf_dhistogram(Fingers, ~Sex.resid)") %>%
check_function("gf_dhistogram") %>% {
check_arg(., "object") %>% check_equal()
check_arg(., "gformula") %>% check_equal()
}
)
ex() %>% check_function("gf_facet_grid") %>% {
check_arg(., "...") %>% check_equal(incorrect_msg = "Make sure you keep the code to create a grid faceted by `Sex`")
}
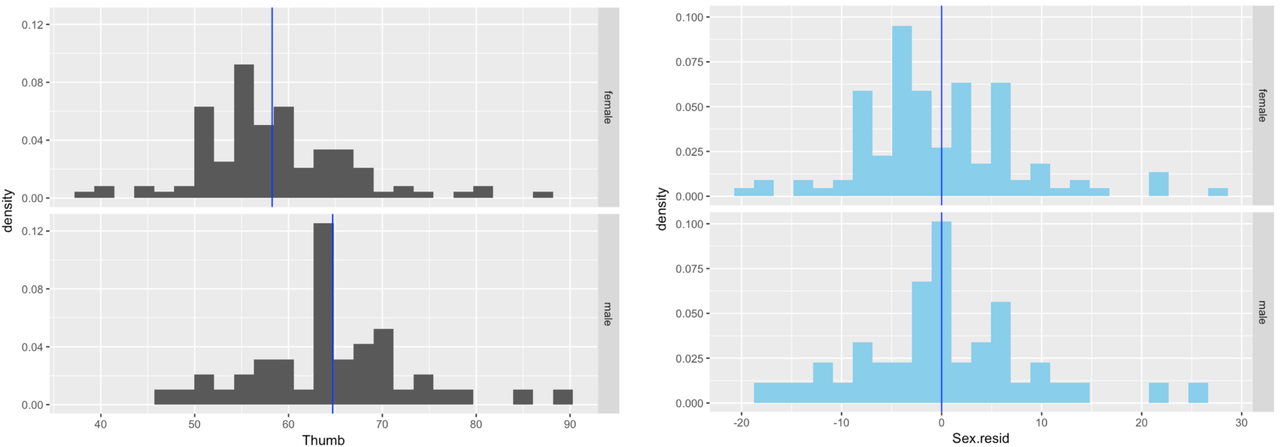
In the activity below, we’ve depicted the density histograms of Thumb by Sex (in black) next to the histograms of Sex.resid by Sex (in skyblue).
We can add the means of Thumb for females and males to the Thumb histograms with some R code. First, we calculate the mean Thumb length for each Sex group and save it in an R object called Thumb.stats:
Thumb.stats <- favstats(Thumb ~ Sex, data = Fingers)Then we chain on (%>%) a vertical line on our histogram with this code.
gf_vline(xintercept=~mean, data=Thumb.stats)Here we have provided the code to add mean lines for each Sex group to the Thumb histograms. Modify the next chunk of code to add mean lines for each Sex group to the Sex.resid histograms.
require(tidyverse)
require(mosaic)
require(Lock5Data)
require(supernova)
Sex.model <- lm(Fingers$Thumb ~ Fingers$Sex)
Fingers$Sex.resid <- resid(Sex.model)
# this creates histograms with lines at the mean of Thumb length for each Sex
Thumb.stats <- favstats(Thumb ~ Sex, data = Fingers)
gf_dhistogram(~Thumb, data = Fingers) %>%
gf_facet_grid(Sex ~ .) %>%
gf_vline(xintercept = ~mean, color = "blue", data = Thumb.stats)
# modify this code to add lines to represent the mean Sex.resid of each Sex group
Sex.resid.stats <- favstats(Sex.resid ~ Sex, data = Fingers)
gf_dhistogram(~Sex.resid, data = Fingers, fill = "skyblue") %>%
gf_facet_grid(Sex ~ . )
Thumb.stats <- favstats(Thumb ~ Sex, data = Fingers)
gf_dhistogram( ~ Thumb, data = Fingers) %>%
gf_facet_grid(Sex ~ .) %>%
gf_vline(xintercept = ~mean, color = "blue", data = Thumb.stats)
Sex.resid.stats <- favstats(Sex.resid ~ Sex, data = Fingers)
gf_dhistogram( ~ Sex.resid, data = Fingers, fill = "skyblue") %>%
gf_facet_grid(Sex ~ . ) %>%
gf_vline(xintercept = ~mean, color = "blue", data = Sex.resid.stats)
ex() %>% {
check_function(., "gf_vline", index = 1)
check_function(., "gf_vline", index = 2)
check_function(., "gf_facet_grid", index = 1)
check_function(., "gf_facet_grid", index = 2)
check_function(., "gf_dhistogram", index = 1)
check_function(., "gf_dhistogram", index = 2) %>%
check_arg("data") %>% check_equal()
}